Single Cell Analysis
Door Josh Loecker

1. Snakemake Pipeline
1.1. Tools
1.1.1. Alignment: Cell Ranger
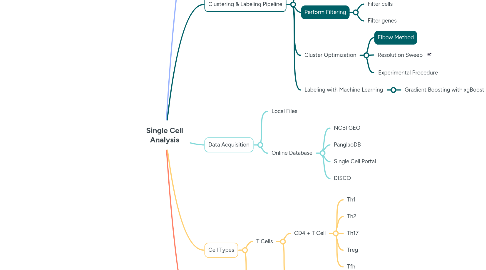
2. Clustering & Labeling Pipeline
2.1. Reading Matrix Files
2.1.1. Directories
2.1.1.1. 10x Matrix
2.1.2. Files
2.1.2.1. HDF
2.1.2.2. 10x h5
2.1.2.3. Matrix Files
2.1.2.4. h5ad
2.2. Perform Filtering
2.2.1. Filter cells
2.2.2. Filter genes
2.3. Cluster Optimization
2.3.1. Elbow Method
2.3.2. Resolution Sweep
2.3.3. Experimental Procedure
2.4. Labeling with Machine Learning
2.4.1. Gradient Boosting with xgBoost
3. Data Acquisition
3.1. Local Files
3.2. Online Database
3.2.1. NCBI GEO
3.2.2. PanglaoDB
3.2.3. Single Cell Portal
3.2.4. DISCO
4. Cell Types
4.1. T Cells
4.1.1. CD4 + T Cell
4.1.1.1. Th1
4.1.1.2. Th2
4.1.1.3. Th17
4.1.1.4. Treg
4.1.1.5. Tfh
4.1.2. CD8+ T Cell
